One approach of decreasing feed volumes but still stoichiometrically supplying nutrients to cells is to concentrate the feed media. The preparation of highly concentrated feed media is, however, challenging because of the limited solubility of medium components. The solubility of many amino acids can be increased by adjusting the pH (3). Recently, surfactants have been used to increase the solubility of chemicals in the feed media (4). In this study, the authors compared strategies including raising the pH and adding surfactant(s) in the preparation of concentrated feed media. In the experiment, the authors fed less volume of concentrated feeds to the cell cultures, as opposed to the constant volume feeding strategy used by Hossler et al. (4). Among the surfactants evaluated, Poloxamer 188 (P188) (i.e., Pluronic F68) is a nonionic surfactant that is an additive in mammalian cell culture media and drug substance formulation (5-7). It was found that the feed volume could be reduced by 2.5-fold with the concentrated media supplemented with higher levels of P188, while volumetric productivity was improved by 10-50%. The improvement of productivities was likely due to the enhancement of protein folding and secretion machinery suggested by quantitative polymerase chain reaction (qPCR) array assay. Supplementing surfactant and raising pH approaches were successfully used in the preparation of concentrated feed media for 7-L and 5000-L bioreactor runs, respectively. These straightforward approaches can be applied to large-scale production of recombinant proteins.
Materials and methods
Cell lines. Three recombinant Chinese hamster ovary (CHO) cell lines, developed in-house to produce three different monoclonal antibodies, were used in this study. Cell lines A and B are dihydrofolate reductase-deficient (DHFR-deficient) DG44 cell lines. Cell line C is a CHO glutamine synthetase (GS) cell line. Frozen vials of the cell lines were thawed and grown in shake flasks containing Bristol-Myers Squibb’s proprietary basal media. The flasks were shaken in incubators at 36.5 °C with 5% CO2.
Media preparation. To prepare feed media for cell lines A and B, the formulated powder was added into water for injection (WFI) or cell-culture-grade water at 37°C, followed by addition of Polysorbate 80 (PS80) or P188. 10 N sodium hydroxide (NaOH) was used to raise the pH until the solution was clear. After the chemicals were completely dissolved, the medium was made up to the required volume by adding WFI or cell-culture-grade water and filtered through a 0.22-µm filter (Millipore). The medium pH was measured by a pH meter (Thermo Scientific), and the medium osmolality was measured by an osmometer (Advanced Instruments). This medium is referenced as M1 in Table I.
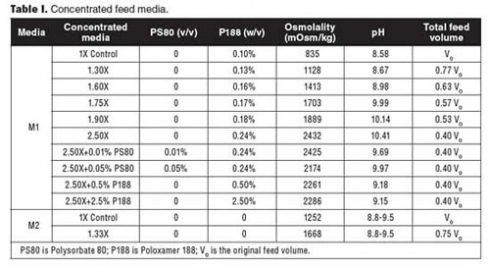
To prepare the feed medium for cell line C, the formulated powders were added into WFI or cell-culture-grade water sequentially at room temperature. 10 N NaOH was used to raise the pH until the solution was clear. After mixing well, the solution was made up to the required volume and filtered through a 0.22 µm filter (Millipore). The pH and osmolality of the medium were measured. This medium is referenced as M2 in Table I.
Fed-batch studies in shake flasks and bioreactors. Fed-batch studies in shake flasks were carried out in 250-mL Erlenmeyer flasks in incubators at 36.5 oC with 5% CO2. Applikon 7-L benchtop bioreactors were used for scaling up shake-flask studies. The process for cell line C was scaled up to the 5000-L production bioreactor.
Cell counts, antibody titer, protein quality analysis, and qPCR array. Cell-culture samples were counted by a Vi-CELL (Beckman-Coulter) using the trypan blue exclusion method. The pH and metabolites in spent media were measured by a NOVA FLEX instrument (Nova Biomedical). Antibody titer was measured using high-performance liquid chromatography (HPLC) (Waters Corporation) equipped with a protein A column (Applied Biosystems). The protein charge profile was analyzed by imaged capillary isoelectric focusing (iCIEF) (ProteinSimple, Santa Clara, California). Antibody N-glycosylated forms were analyzed by PNGase F digestion, reductive amination labeling, followed by separation using ultra-performance liquid chromatography (UPLC).
In a qPCR array, 3 × 106 cells of day 9 and day 11 from each treatment were used to extract messenger ribonucleic acid (mRNA) following the protocol of RNeasy Mini Kit (Qiagen). 400 ng of total RNA for each sample was reverse transcribed into complementary deoxyribonucleic acid (cDNA) using RT2 First Strand Kit (Qiagen). Four experimental cocktail solutions composed of cDNA synthesis reaction and SABiosciences RT2 qPCR master Mix (Qiagen) were dispensed into 4 × 96 wells (10 µL/well) in 384-well PCR unfolded protein response array (Qiagen). The array was analyzed by ViiA 7 real-time PCR system (Life Technologies). The raw CT values were further analyzed by free web-based RT2 profiler PCR array data analysis version 3.5 (Qiagen). The ratios of gene levels between day 9 and day 11 for each treatment were reported. The gene functions were obtained from the website of the National Center for Biotechnology Information (www.ncbi.nlm.nih.gov/gene).