Optimization of each phase in a chromatographic cycle has a positive impact on productivity.
By Anna Grönberg
 An important area of focus for the biopharma industry is the elucidation and development of robust, efficient, and economic purification strategies for new monoclonal antibodies (mAbs) coming through the pipeline. Since the first therapeutic antibody came onto the market in 1986, up to the end of 2014, 47 products have been approved in the United States or Europe for a number of diseases including various types of cancer, autoimmune, autoinflammatory diseases, and rare diseases (1, 2). A study by BioProcess Technology Consultants estimated that approximately 70 mAb products could be on the market by 2020, which could be associated with up to £125 billion in sales (1).
An important area of focus for the biopharma industry is the elucidation and development of robust, efficient, and economic purification strategies for new monoclonal antibodies (mAbs) coming through the pipeline. Since the first therapeutic antibody came onto the market in 1986, up to the end of 2014, 47 products have been approved in the United States or Europe for a number of diseases including various types of cancer, autoimmune, autoinflammatory diseases, and rare diseases (1, 2). A study by BioProcess Technology Consultants estimated that approximately 70 mAb products could be on the market by 2020, which could be associated with up to £125 billion in sales (1).
Purifying these biological products once produced by cells requires a reliable and optimized chromatography process. An advantage in the area of mAbs is the possibility of using a purification platform approach. The typical platform process is generally comprised of an initial capture step with Protein A-chromatography, a first polishing step with cation-exchange media, and a final polishing step using anion-exchange or multimodal media. Even this platform process requires some fine-tuning and adjustment for new mAb targets, especially at the first polishing step with cation exchange chromatography in bind-and-elute mode. This step often requires more optimization than initial capture and/or final polishing and aims to achieve high-dynamic binding capacity and maximum aggregate and host-cell protein (HCP) clearance. In addition to assessing factors and conditions for the optimization of the purification steps, the quality-by-design (QbD) initiative introduced by FDA requires a deeper understanding of critical process parameters to be obtained during process development. This deeper level of understanding could be directly addressed by increasing the experimental space using high-throughput process development (HTPD).
Leveraging the power of HTPD and DoE for chromatographic optimization
The goal in HTPD is to evaluate numerous factors or conditions in parallel using micro-scale formats. It reduces experimental time frames from weeks to hours, requires low sample consumption, and increases process understanding by exploring a larger experimental window. With HTPD, the typical workflow follows a series of steps, including planning an experimental setup using a systematic approach, carrying out experiments by testing various experimental conditions in a parallel manner, analyzing data using high-throughput analysis, and evaluating data and analyzing the results.
Design of experiments (DoE) provides a statistics-based methodology to identify and define factors affecting the overall process or end product. By studying multiple factors simultaneously, DoE allows a maximum amount of information from the minimum number of experiments. Objectives such as screening, optimization, and robustness testing are set before the studies begin. Data generated by DoE can be used to create models that can apply statistics to predict future responses of a particular system to conditions that so far have not been tested. An example illustrating the use of HTPD and DoE to optimize the first polishing step for a mAb of interest is discussed herein, and the roles of both HTPD 96-well filter plates and mini-columns are highlighted (3).
Case study: Accelerating mAb purification via HTPDThe mAb used in the following case study was produced in Chinese hamster ovary (CHO) cells, and the initial mAb capture step was carried out with an alkali-stabilized, Protein A-derived affinity medium using a standard protocol. An HTPD approach was then taken for the subsequent purification using a high-capacity cation exchanger. First, static binding capacity (SBC) was determined, using 96-well filter plates to allow high-throughput screening of a large number of experimental conditions. 96-well filter plates prefilled with chromatography medium (Cytiva) were run in batch mode. In HTPD, simple absorbance measures such as A280 measurements can be used, and additional analyses carried out for the most critical conditions. In this mAb case study, SBC increased by increasing pH from 4.5 to 6.0 and the salt concentration from 0 to 50 mM NaCl (see Figure 1). Further investigation of binding capacity and selectivity was limited to the known pH limits for the stability of the mAb being tested.
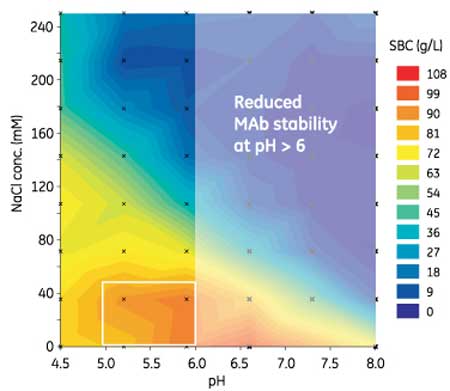 Figure 1: Results of static binding capacity (SBC) screening are displayed in a contour plot, as determined using a high-throughput process development (HTPD) 96-well plate prefilled with a strong cation-exchange chromatography medium.
Figure 1: Results of static binding capacity (SBC) screening are displayed in a contour plot, as determined using a high-throughput process development (HTPD) 96-well plate prefilled with a strong cation-exchange chromatography medium.
Dynamic binding capacity (DBC) was assessed using high-throughput modular mini-columns (Cytiva). By using mini-columns operated in parallel, a larger number of different experimental conditions, such as binding, and/or wash and elution conditions, can be explored in a shorter time frame compared with when using small-scale columns, which are operated in a sequential mode. In this study, a robotic liquid handling system was used to run eight columns, packed with the cationic exchange medium, in parallel. Using a DoE approach, different pH and NaCl concentrations were studied to determine the dynamic binding capacities at selected conditions. A mAb eluate from the Protein A capture step conditioned to the different pH and salt concentrations was loaded onto the columns. Protein concentration in flowthrough fractions were analyzed by UV spectroscopy. DBC was found to increase with pH and NaCl concentration within the explored space, which correlated well with the SBC experiment (see Figure 2).
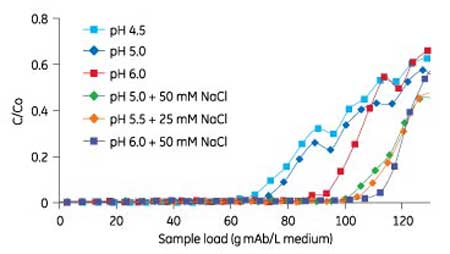 Figure 2: Results of different binding conditions at different pH and NaCl concentrations are shown as an overlay of breakthrough curves generated on PreDictor RoboColumn units (Cytiva). The breakthrough (breakthrough concentration [C]/start concentration [C0]) is plotted against the sample load in gram mAb/Liter medium.
Figure 2: Results of different binding conditions at different pH and NaCl concentrations are shown as an overlay of breakthrough curves generated on PreDictor RoboColumn units (Cytiva). The breakthrough (breakthrough concentration [C]/start concentration [C0]) is plotted against the sample load in gram mAb/Liter medium.
Selectivity between mAb monomer and aggregates was also studied using DoE, analyzing factors such as pH and NaCl concentration. For this purpose, small-scale columns were employed. Aggregate concentration and host-cell protein (HCP) levels in the eluate at a 90% monomer yield were observed. The loading pH was found to have a significant effect on aggregate concentration, where lower pH was optimal for a low level of aggregates. A 2–4 fold reduction in HCP was observed following the cation exchange polishing step (see Table I).
Table I: Effects of different binding and elution conditions on monoclonal antibody (mAb) aggregate and host-cell protein (HCP) content at 90% monomer yield. The center point of the factors is halfway between the lowest and highest levels (pH=5.5, NaCl=25 mM).
The result of process optimization was a DBC of >100 mg mAb/mL medium at a loading condition of pH 5.0 and 50 mM NaCl. A Tricorn 5/100 column (Cytiva) was used to verify the DBC at 10% breakthrough (QB10), which was 109 mg mAb/mL, comparable to the mini-columns used (108 mg mAb/mL). With a load of 50 mg mAb/mL, less than 1% aggregate content was found and monomer yield was above 90%. Even at a higher sample load of 76 mg mAb/mL, 0.9% aggregate at a monomer yield of 93% was achieved, with an HCP concentration of 170 ppm and less than 1 ppm of leached Protein A from the capture step (Figure 3).
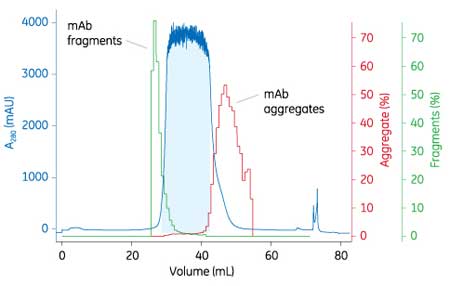 Figure 3: Selectivity between monoclonal antibody (mAb) monomers and aggregates at a higher sample load following the optimized first polishing step in a lab-scale column, showing the percentage of fragments and aggregates in the resulting fractions. A280=absorbance at 280 nm wavelength as measured with UV-visible spectroscopy; mAU=1/1000th of an absorbance unit as measured with UV-visible spectroscopy.
Figure 3: Selectivity between monoclonal antibody (mAb) monomers and aggregates at a higher sample load following the optimized first polishing step in a lab-scale column, showing the percentage of fragments and aggregates in the resulting fractions. A280=absorbance at 280 nm wavelength as measured with UV-visible spectroscopy; mAU=1/1000th of an absorbance unit as measured with UV-visible spectroscopy.
Considerations when using mini-columns on a robotic liquid-handling station
The slowest flow rate when using a 1-mL syringe on the TECAN Freedom Evo 150 robot, for example, is 1 µL/s. With PreDictor RoboColumn units 200 µL (Cytiva), this flow rate will result in a residence time (RT) of 3.33 minutes. Using the larger column volumes (i.e., 600 µL) will increase the RT to 10.0 min with the slowest flow rate. Thus, depending on what residence time that is needed, the relevant column volume should be selected.
To avoid system liquid dilution of the protein sample during sample injection to PreDictor RoboColumn units, an excess volume of 400 µL protein sample needs to be aspirated. This means that with the 1000-µL syringe, 1000 µL of protein sample can be aspirated, but only 600 µL will be dispensed into the columns. The issues with dilution have not been observed for buffers and salt containing aqueous solutions.
Optimizing binding capacity
The dynamic binding capacity is dependent upon several experimental parameters (e.g., the charge and size of the target molecule, pore size, and the charge of the chromatography media, as well as the ionic strength of the loading buffer). The highest DBC on an ion-exchange chromatography media is typically expected at low conductivity and at a pH far away from the isoelectric point (pI), where the protein charge is high. For some proteins, increasing conductivity and decreasing protein charge can improve capacity (4, 5). Careful investigation of the effect of pH and salt concentration on dynamic binding capacity should be undertaken in order to find the highest binding capacity.
Productivity
The productivity, expressed as amount of product per unit of time and unit of chromatography media, is determined by the breakthrough capacity of the chromatography media, the recovery of active product, and the cycle time.
The cycle time can be reduced by selecting modern chromatography media that can be operated under high volumetric flow rates. Even though the binding phase may be constrained to a certain residence time due to mass transfer limitations and a decrease in capacity at higher flow velocities, equilibration, wash, and elution can be carried out at maximum allowed operating velocity.
Optimization of each phase in a chromatographic cycle to decrease the cycle time will have a positive impact on productivity. By transferring a continuous gradient elution to a step gradient elution, for example, there is a significant decrease in elution volume, which shortens cycle time and increases productivity.
Conclusion
The use of a HTPD-based workflow in conjunction with DoE has been described as a method to optimize conditions for the first polishing step of mAb purification. This workflow is compatible with robotic liquid handling stations for 96-well filter plates and mini-columns and steps can be taken to reduce the cycle time while increasing productivity, including improved recovery of active products.
Plates and mini-columns, however, have different advantages and limitations for various applications. When used in combination, they provide a powerful platform for quickly and efficiently determining which conditions work best for a particular mAb of interest, with excellent correlation and reproducibility at larger scale. Filter plates are useful tools to screen for a broad range of conditions, whereas mini-columns can be used to investigate dynamic parameters. Lab-scale columns can then be used to follow-up from the high-throughput systems to determine selectivity and verify binding conditions.
References
1. D.M. Ecker, S.D. Jones, and H.L. Levine, mAbs 7 (1), pp. 9-14 (January/February 2015).
2. A. Beck et al., Nature Rev. Immunol. 10, pp. 345-352 (May 2010).
3. Cytiva, “Optimization of dynamic binding capacity and aggregate clearance in a monoclonal antibody polishing step,” Application note 29145068 AA (2015).
4. C. Harinarayan et al., Biotechnol. Bioeng. 95 (5), pp. 775-787 (2006).
5. Cytiva “Screening and optimization of the loading conditions on Capto S.,” Application note 28-4078-16 AA (2006).
About the Author
Anna Grönberg is staff research engineer, R&D, at Cytiva.
ALL FIGURES ARE COURTESY OF THE AUTHOR.